- Lesson 9: Data Query in a Federated Environment
- Slides from the video
- why do we need distributed systems
- hQuery
- Informatics for Integrating Biology and the Bedside (i2b2)
- PopMedNet
- Query Health
- Recap
- Supplementary resources
Lesson 9: Data Query in a Federated Environment
- Distributed query: securely obtaining useful data from diverse EHR systems and other sources.
- ONC supported Query Health initiative.
- Query Health initiative related video (The first 18 minutes is a good general introduction to distributed query that you should watch. The rest describes individual query reporting systems.)
Slides from the video


- Govern is more important


why do we need distributed systems
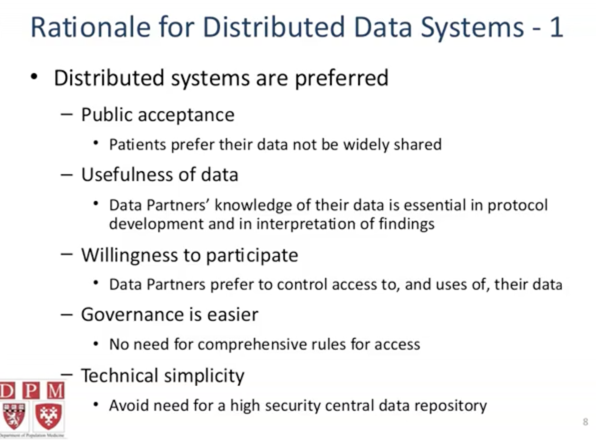
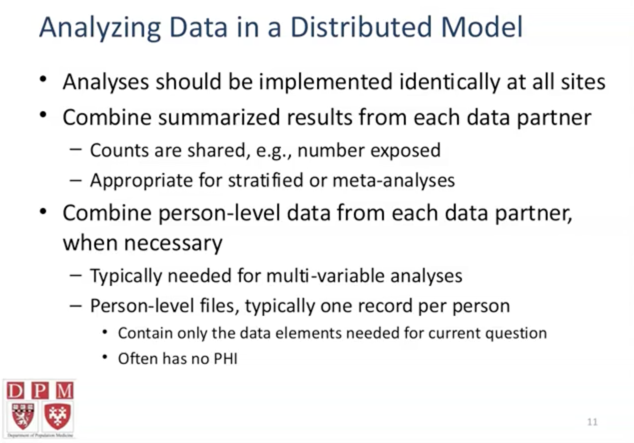






hQuery
- hQuery’s goal: simplicity. limited but well standardized data set (in quality measurement and other common clinical activities).
- point-and-click interface (called the hQuery composer) to non-technical healthcare providers.
- the possible queries are somewhat limited.
- hQuery uses a simple patient information model to facilitate query building by nontechnical users.
- hQuery Gateway
- located at source system
- receives queries in a standard format
- forwards queries to an adapter
- the adapter translates the query, receives response and converts it into the standard data model for transmission back.
- Video about hQuery.
- HQuery presents results in an attractive way (frequency, time and geographic distributions).

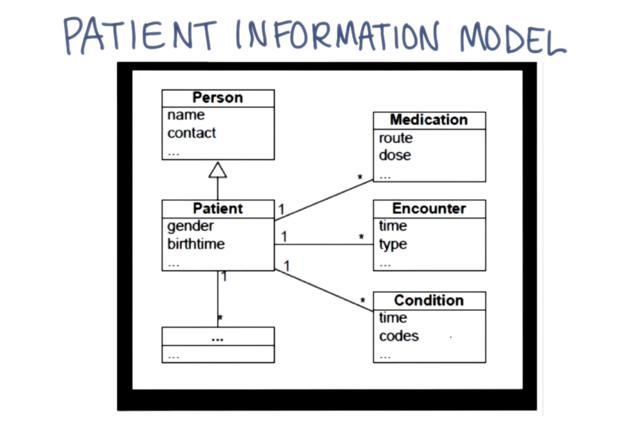
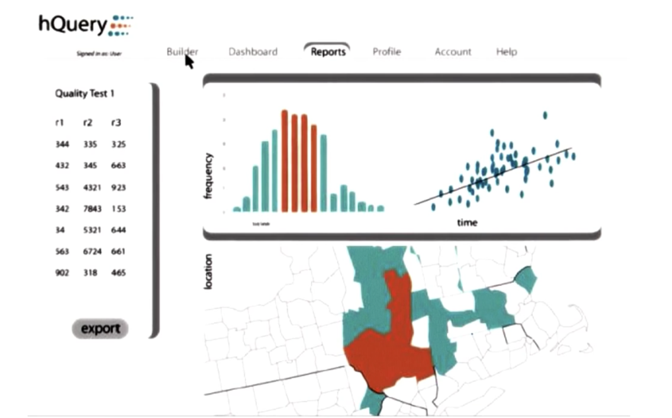

- the curly braces problem is a technical sub-issue of interoperability problem
Informatics for Integrating Biology and the Bedside (i2b2)

- is NIH-funded effort based at Partners HealthCare System in Boston, Mass. * mission: enable clinical investigators to conduct research using genomics and biomedical informatics.
- is sophisticated to support complex research environments
- provides an easily understood database schema
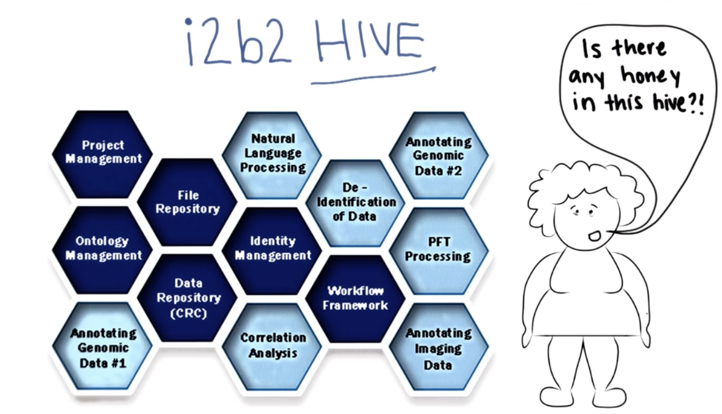
- i2b2 “cells” communicate via web services and are together called a “hive”.
- Custom cells can be added to the hive.
- i2b2 users group (share information and applications; create the potential for translational research)

- extracting clinical concepts or features from free text format clinical notes
PopMedNet
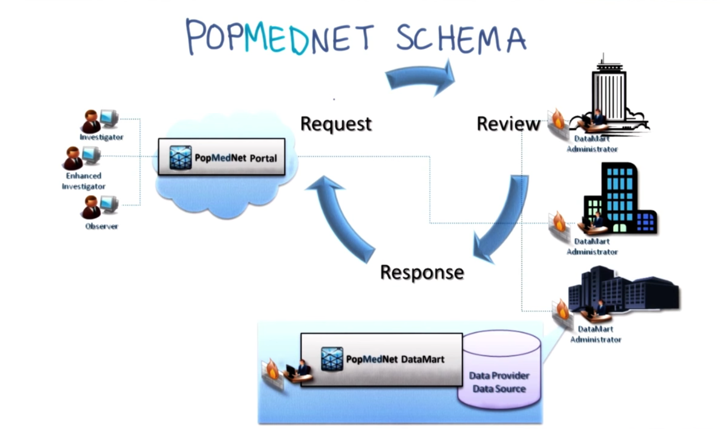
- each source to maintain control of its data (sometimes referred to as “data lockers”)
- be capable of accepting and responding to queries.
- PopMedNet is intended to support medical product safety analysis



Query Health
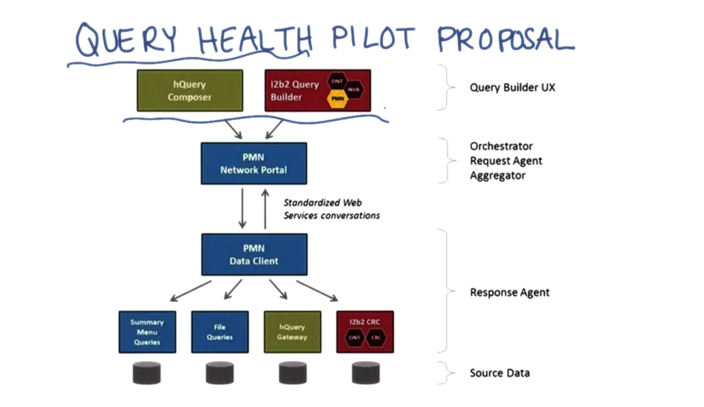
distributed query standards.
Four kinds of standards:
- Envelope standards: define the package for sending/receiving queries
- Format standards: provide a Declarative specification of the query
- Results standards (such as QRDA): specify the format and packaging of the query results
- Data Model standards (such as CEDD common data element definitions): specify the data model that will link data from the contributing systems.


- Declarative specification: expresses the desired result.
- Procedural specification: provides a recipe for achieving the goal
- E.g. 1 Curly braces problem with the Arden syntax. Arden provides a declarative standard but the procedure will be different depending on the system that is the source of the data.
- e.g. 2 The hQuery Gateway receives a standard query on one side but knows how to get it done on the other.

Query envelope standard
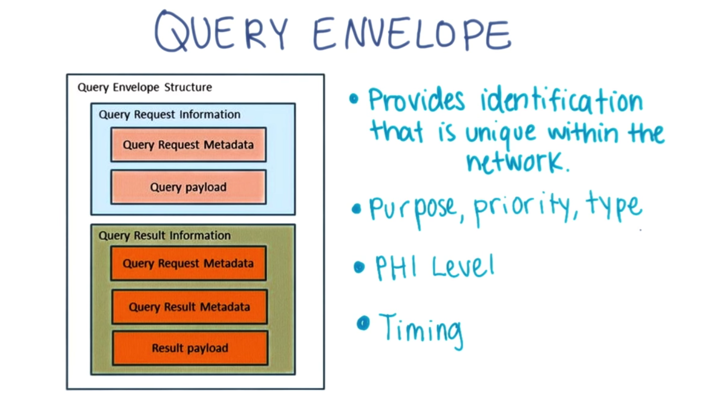
The Query Envelope standard serves to provide
- identification that is unique within the network;
- the information requestor ID including name, email and organization;
- the purpose, priority and type of the query
- 7 purpose codes (e.g. TREAT for treatment),
- its Priority from 1-5 (1 highest),
- its type Type, 1-20 characters
- PHI level (Aggregated, Limited, De-identified, PHI)
- its Timing, Submission date/time and optional execution date/time.
Query Format
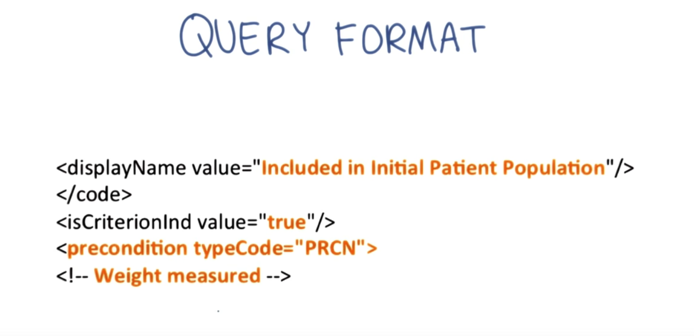


- the Health Quality Measure Format (HQMF): an HL7 standard format for documenting the content and structure of a quality measure in an XML document format based on the HL7 RIM.
- It can consist of three levels of detail:
- metadata: who wrote it, the dates over which it is valid, who validated it, and other details about how the measure works or is used;
- human narrative: measure description, data criteria, measure population and measure observations;
- computer instructions: how to count and compute the results of the measure.
- The example XML provides instructions to the computer to only include patients who have had a weight measured, presumably in the numerator of a weight screening quality metric that, as you should recall from Lesson 2, is a requirement of Meaningful Use.
the standard for query results
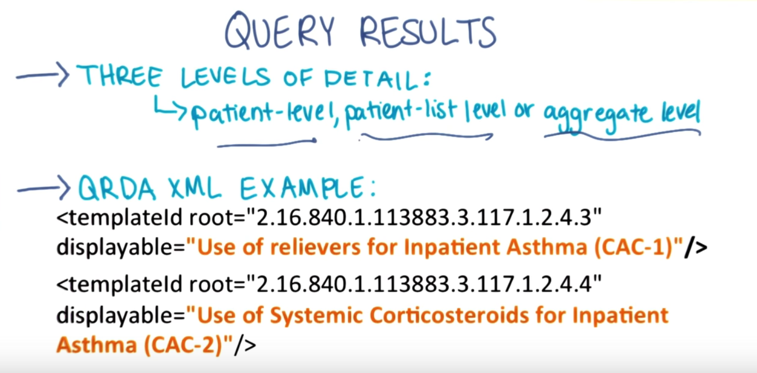

- QRDA in Lesson 8. Results can be reported at three levels of detail: patient-level, patient-list level or aggregate level.
- Example: the two result templates refer to NQF defined quality measures for inpatient asthma care.
Query Data Dictionary
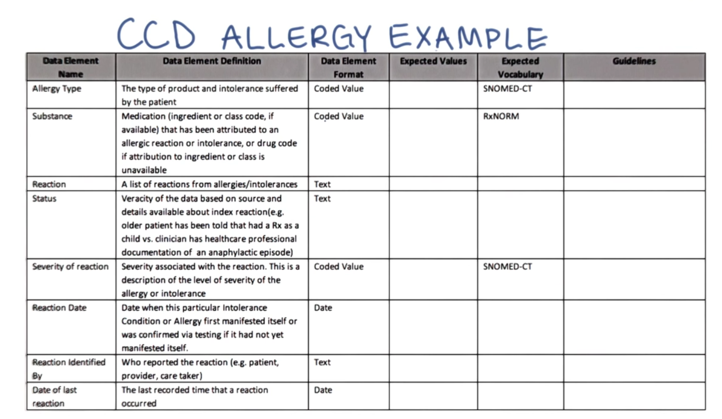
- The Clinical Element Data Dictionary (CEDD): use in setting up source data in support of distributed queries within a larger Query Health solution.
- it is not intended as a new standards development effort and began with the elements already specified in the CCD and which most EMRs already support
- Here’s a simple example for the patient allergy data element showing how it ties to already accepted standards used in the CCD.

Recap
In this lesson we
- examined how data from multiple EHR systems can be queried and aggregated for diverse purposes.
- e.g. quality reporting and advanced clinical research.
- All of these technologies provide a framework over the many non-interoperable EHR systems.
- Different technological solutions each of which is optimized for a specific problem in a specific cross-institutional context.
Supplementary resources
Key Concepts/Vocabulary
●Distributed query ● Distributed query standards ● Data warehouse ● Data lockers ● hQuery ● i2b2 ● PopMedNet ● Query Health ● Health Quality Measure Format (HQMF) ● Quality Reporting Document Architecture (QRDA)
Readings
Videos
Graphics
2015-10-19 初稿